readr::read_lines("./data/murders.csv", n_max = 3) ## there is a header[1] "state,abb,region,population,total" "Alabama,AL,South,4779736,135"
[3] "Alaska,AK,West,710231,19" MATH/COSC 3570 Introduction to Data Science
| Function | Format | Typical suffix |
|---|---|---|
read_table() |
white space separated values | txt |
read_csv() |
comma separated values | csv |
read_csv2() |
semicolon separated values | csv |
read_tsv() |
tab delimited separated values | tsv |
read_fwf() |
fixed width files | txt |
read_delim() |
general text file format, must define delimiter | txt |
Be careful: The suffix usually tells us what type of file it is, but no guarantee that these always match.
readr::read_lines("./data/murders.csv", n_max = 3) ## there is a header[1] "state,abb,region,population,total" "Alabama,AL,South,4779736,135"
[3] "Alaska,AK,West,710231,19" read_csv() prints out a column specification giving us delimiter, name and type of each column.
murders_csv <- read_csv(file = "./data/murders.csv")
# Rows: 51 Columns: 5
# ── Column specification ─────────────
# Delimiter: ","
# chr (3): state, abb, region
# dbl (2): population, total
head(murders_csv)# A tibble: 6 × 5
state abb region population total
<chr> <chr> <chr> <dbl> <dbl>
1 Alabama AL South 4779736 135
2 Alaska AK West 710231 19
3 Arizona AZ West 6392017 232
4 Arkansas AR South 2915918 93
5 California CA West 37253956 1257
6 Colorado CO West 5029196 65## View data in RStudio
view(murders_csv)Which type is the column vector x? Why?
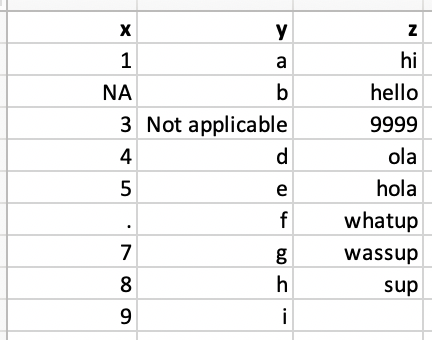
read_csv("./data/df-na.csv")# A tibble: 9 × 3
x y z
<chr> <chr> <chr>
1 1 a hi
2 <NA> b hello
3 3 Not applicable 9999
4 4 d ola
5 5 e hola
6 . f whatup
7 7 g wassup
8 8 h sup
9 9 i <NA> read_csv() only recognizes ” “ and NA as a missing value.na.read_csv("./data/df-na.csv",
na = c("", "NA", ".", "9999", "Not applicable"))
# A tibble: 9 × 3
x y z
<dbl> <chr> <chr>
1 1 a hi
2 NA b hello
3 3 <NA> <NA>
4 4 d ola
5 5 e hola
6 NA f whatup
7 7 g wassup
8 8 h sup
9 9 i <NA> read_csv("./data/df-na.csv",
col_types = cols(col_double(),
col_character(),
col_character()))Warning: One or more parsing issues, call `problems()` on your data frame for details,
e.g.:
dat <- vroom(...)
problems(dat)# A tibble: 9 × 3
x y z
<dbl> <chr> <chr>
1 1 a hi
2 NA b hello
3 3 Not applicable 9999
4 4 d ola
5 5 e hola
6 NA f whatup
7 7 g wassup
8 8 h sup
9 9 i <NA> problems()
# A tibble: 1 × 5
# row col expected actual file
# <int> <int> <chr> <chr> <chr>
# 1 7 1 a double . "" | type function | data type |
|---|---|
col_character() |
character |
col_date() |
date |
col_datetime() |
POSIXct (date-time) |
col_double() |
double (numeric) |
col_factor() |
factor |
col_guess() |
let readr guess (default) |
col_integer() |
integer |
col_logical() |
logical |
col_number() |
numbers mixed with non-number characters |
col_numeric() |
double or integer |
col_skip() |
do not read |
col_time() |
time |
head(iris, n = 3) Sepal.Length Sepal.Width Petal.Length Petal.Width Species
1 5.1 3.5 1.4 0.2 setosa
2 4.9 3.0 1.4 0.2 setosa
3 4.7 3.2 1.3 0.2 setosa## save iris data to "./data/iris.csv"
iris |> write_csv(file = "./data/iris.csv")
df_iris <- read_csv(file = "./data/iris.csv")
df_iris# A tibble: 150 × 5
Sepal.Length Sepal.Width Petal.Length Petal.Width Species
<dbl> <dbl> <dbl> <dbl> <chr>
1 5.1 3.5 1.4 0.2 setosa
2 4.9 3 1.4 0.2 setosa
3 4.7 3.2 1.3 0.2 setosa
4 4.6 3.1 1.5 0.2 setosa
5 5 3.6 1.4 0.2 setosa
6 5.4 3.9 1.7 0.4 setosa
7 4.6 3.4 1.4 0.3 setosa
8 5 3.4 1.5 0.2 setosa
# ℹ 142 more rows10-Import Data
tidyverse package.In lab.qmd ## Lab 10 section,
read_csv() and call it ssa_male.Age (x-axis) vs. LifeExp (y-axis). The type is “line”, and the line color is blue. Add x-label, y-label and title to your plot.penguins_raw in the R palmerpenguins package to “./data/penguins_df.csv”. Print penguins_df.csv out.| Function | Format | Typical suffix |
|---|---|---|
read_excel() |
auto detect the format | xls, xlsx |
read_xls() |
original format | xls |
read_xlsx() |
new format | xlsx |
excel_sheets() gives us the names of all the sheets in an Excel file.library(readxl)
excel_sheets("./data/2010_bigfive_regents.xls")[1] "Sheet1" "Sheet2" "Sheet3"sheet argument to read sheets other than the first.excel_sheets("./data/2010_bigfive_regents.xls")[1] "Sheet1" "Sheet2" "Sheet3"(data_xls <- read_xls(path = "./data/2010_bigfive_regents.xls",
sheet = "Sheet3",
skip = 1))# A tibble: 19 × 6
Scores `131024` `113804` `104201` `103886` `91756`
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 10 NA 64 8 227 34
2 11 6 83 11 217 58
3 12 23 87 7 28 67
4 13 1 54 16 230 42
5 14 3 145 18 303 57
6 15 58 151 50 192 98
7 16 1 129 13 156 125
8 17 73 214 59 163 115
# ℹ 11 more rowspd.read_csv
pd.DataFrame.to_csv
pd.read_csvpd.DataFrame.to_csv Unnamed: 0 x y
0 0 1 a
1 1 2 b
2 2 3 c10-Import Data
>I have a R object that cannot be saved as a csv or excel file, for example, a list. How can I save that object, and reuse it later? Please teach me with a simple example.